|
|
Post by djoser-xyyman on Jan 5, 2018 11:47:06 GMT -5
Ancient or modern Egypt. Modern European lineage is NOT found in Egypt. mtDNA H1 and H3 are over 20,000 years old. U5 maybe close to 35,000years old. Ancient Egyptians(and Modern) Egyptians has absolutely no relation to Egypt past or present. Nein!!!   |
|
|
|
Post by zarahan on Jan 5, 2018 16:50:53 GMT -5
First, If you look at the charts Abusir do NOT cluster with Europeans. Second, look at the “Eurasians” they cluster with. It is NOT Syrians, Druze, Iraqi’s, Iranians, Jordanians etc. The “Eurasians” they cluster closest with are Bedouins and Yemenis NOT the usual suspects you consider “middle Easterners”. The African populations of the Great Lakes were NOT included in the study of the Abusir. Instead, they used the distant YRIIndeed. And their omission of the Great Lakes which in previous studies showed some links to Egypt is telling. Jumping to the distant YRI far to the south and West is typical. HOw many studies have we seen in the past that sample far to the north in Egypt, while excluding the "darker" historic south? They have been running that game in cranial and dental studies, etc and the same pattern is used in DNA studies. 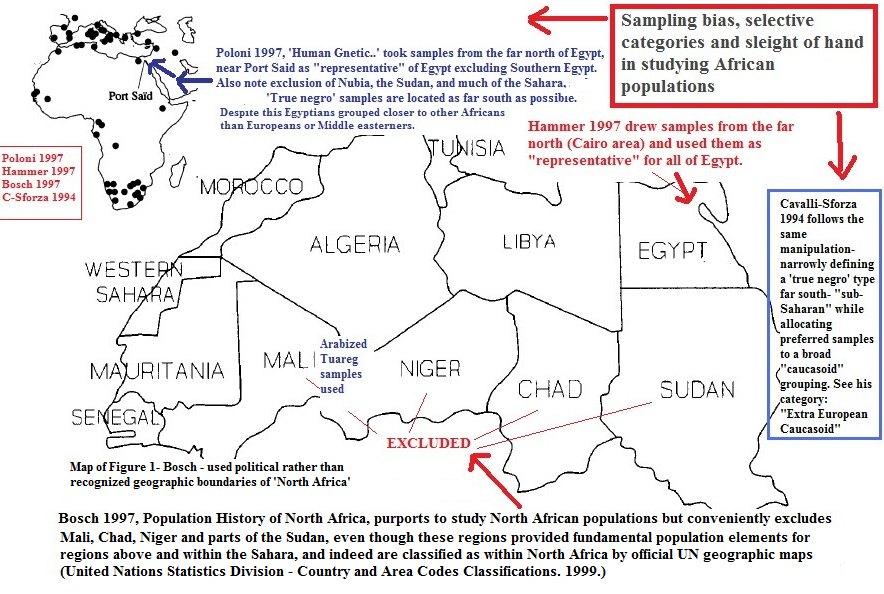  Coastal North African sampling issues/bias 
|
|
|
|
Post by zarahan on Jan 5, 2018 17:12:58 GMT -5
How would you link the earlier DNATribes findings re Great Lakes links
to the above that you post?
|
|
|
|
Post by melanitex on Jan 5, 2018 17:54:03 GMT -5
What is incorrect? What physical proof or data do you have that suggest Europeans were present in AE? Do you understand the data of the Abusir mummies. I posted the charts from the study. Do you understand it? The Abusir paper did NOT state the Abusir mummies were Europeans it said they cluster with “Eurasians”. Yes, There is a difference!!! First, If you look at the charts Abusir do NOT cluster with Europeans. Second, look at the “Eurasians” they cluster with. It is NOT Syrians, Druze, Iraqi’s, Iranians, Jordanians etc. The “Eurasians” they cluster closest with are Bedouins and Yemenis NOT the usual suspects you consider “middle Easterners”. The African populations of the Great Lakes were NOT included in the study of the Abusir. Instead, they used the distant YRI. Also they did not disclose STR profile which was deliberate deception by the researchers. Listen, I don’t want to talk down at you but, really, you are not equipped to have this discussion/conversation with me. If you read my posting or the paper you would not be asking those questions. Oh right my bad....I accidentally misunderstood the meaning of Eurasian..... |
|
|
|
Post by kel on Jan 6, 2018 21:36:42 GMT -5
'Oh right my bad....I accidentally misunderstood the meaning of Eurasian....'
It wasnt an accident at all.....
Its just years of dishonest programming by academics have had its effect.
The whole point of the Eurasian thing is to fool people into consciously and subconsciously equating that with European.
|
|
|
|
Post by djoser-xyyman on Jan 9, 2018 8:22:58 GMT -5
That is exactly what it is, programming. Eurasian=Caucasian. European has no genetic affinity to AEians. None! Look at the Abusir Charts. They used the label "Eurasians" to deceive into believing AEians are Europeans. But let's take it a step further. Forget the game they played when they left out Great Lakes Africans when in the past papers(like JAMA the Amarna's) these were closest to AEians. Look at the "Eurasian" population Abusir mummies are closest to base upon autosomes. And Look at also their uniparental markers. Abusir do NOT clusters with Syrians, Iraqi, Druze, Jordanians, Iranians, Kuwaiti etc. Population you visualize as "middle Eastern". Understand it is a game!! Abusir cluster with Bedoiuns and Yemenis. Two heavily Africanized "Near East" populations. And if you take the Abusir uniparental(mtDNA) markers into consideration they are Africans from up the Great Lakes. Like Tanzania, Ugandan, Ethiopia and so on. The very same population that was NOT included in the study. Understand the game. Instead they chose the distant Yoruba as SSA. Tricksters!!
|
|
|
|
Post by djoser-xyyman on Jan 10, 2018 8:30:56 GMT -5
Goodness we have the internet and we can go into databases and manually do a compare. As a result we can't be easily fooled. Of course they are keeping data on lock down. But with the little they release and the tools available we can do a hands on painful tedious analysis without their input. It would be nice to have their full data but they would not allow it. Because their lies will be quickly exposed like with the Amarna's. And DNATribes as reputable company they would have probably given the data to are no longer in that business since the founder met an untimely death.
|
|
|
|
Post by zarahan on Jan 12, 2018 15:41:00 GMT -5
What other studies besides the DNAtribes one shows the Great-Lakes AE links?
|
|
|
|
Post by djoser-xyyman on Jan 12, 2018 20:45:05 GMT -5
  YOU and anyone can do it themselves. The autosomal STR profile of the Amarnas is freely available from the JAMA report. Software is freely available on the web. |
|
|
|
Post by africurious on Jan 13, 2018 0:37:28 GMT -5
^In other words none, lol.
Zarahan I’d suggest you be careful of the stock you put into dna tribes Amarna article. They’re not a scientific journal, just a for profit genetic testing company for the layman.
As far as their Amarna results:
1. The MLI score they use to assign regional ancestry is a proprietary formula and isn’t an academic standard
2. The MLI scores for the Great Lakes weren’t particularly high. Most of the scores were in the low hundreds. Note there’re people who’ve used dna tribes for ancestry test and get scores in the hundreds of thousands. Likely scenario is that due to dna tribes reference sample the best interpretation is that the Amarna’s scores are indicating African ancestry generally. The reference sample isn’t good enough to get a more detailed pinpoint in Africa. Great Lakes, southern Af and West Af scored really high but the regions closer to AE such as the Sahel and the horn scored much lower. Does that even make intuitive sense? They scored lower because in current times these 2 regions have higher non-african admixture than other regions.
|
|
|
|
Post by djoser-xyyman on Jan 13, 2018 5:51:01 GMT -5
REPEAT!!!!!!   YOU and anyone can do it themselves. The autosomal STR profile of the Amarnas is freely available from the JAMA report. Software is freely available on the web. Read more: egyptsearchreloaded.proboards.com/thread/2611/europeans-any-connection-ancient-egypt?page=2#ixzz543nlkI1RWhat is wrong with you people. Why is it so difficult to understand 1. DNATribes did NOT perform any study. They took public available data from JAMA and ran the analysis using their database of samples. THEY DID NOT DO ANY TESTING!!! 2. YOU and or Joe Public can do the same analysis. 3. Saying DNAtribe is a "for profit" company shows you do not understand what is going on or is deliberately confusing the issue. 4. DNAConsultants gave similar results. Do you know that? Stop confusing readers if you do not know what you are talking about |
|