Post by djoser-xyyman on Dec 20, 2018 12:37:24 GMT -5
These Europeans never cease to baffle me with their logic and their lies. Are they that obtuse. Their abstract states Khoe-San got their light skin from an imaginary Near East population but the admit LWK/Khoi_San has MORE DIVERSE SLC24A5 and it is also much much older in Africa!!!! I am reading the paper looking at the data but I am baffle with their conclusion without providing comparable data. God Damn! These white people are crazy and manic.
-----------------
Rapid evolution of a skin-lightening allele in southern African KhoeSan - Meng Lina, Carlos D. Bustamante and Brenna M. Henna
Quote:
“e.g., ∼50% lighter than equatorial Ghanaians. The KhoeSan form the earliest divergence among modern human lineages, branching off from other populations as early as 100 kya to
150 kya. This raises the possibility that the light to intermediate skin of KhoeSan observed today **reflects the ancestral state of the phenotype** and light skin pigmentation in Eurasians
evolved from standing genetic variation within southern Africa .”
“SLC24A5 is among the top signals in positive selection scans of Europeans and Ethiopians, and has long been recognized to play a role in pigmentation pathways.
The nonsynonymous p.Ala111Thr mutation (G → A) at rs1426654 causes a melanin-reduced phenotype in zebrafish. In humans,
the derived allele has swept to fixation or is present at high frequency in many European, Near Eastern, and South Asian populations ; rs1426654 has also been associated with skin pigmentation differences in admixed populations who have recent European ancestors
(i.e., African Americans) and within South Asians. In contrast, the absence of the derived allele in East Asians and
MOST??? sub-Saharan African populations led to the hypothesis that this mutation originated in Europe, or, potentially, the Near East within
the past 10 kya to 35 kya (Table 1).”
“Among the 430 individuals, the derived allele (rs1426654*A) is present at a high frequency of 32.5% in ≠Khomani
And 53.5% in Nama. Queries of rs1426654 in publicly available datasets show that the allele is absent or [/b]at a very low frequency[/b] in
other populations from western, central, and southern Africa (Fig. 1A and SI Appendix, Table S1). Other populations with a moderate to
high frequency of the allele outside Europe have all experienced gene flow from the Near East or Europe during the Holocene, including
those in northern and eastern Africa, and South Asia”
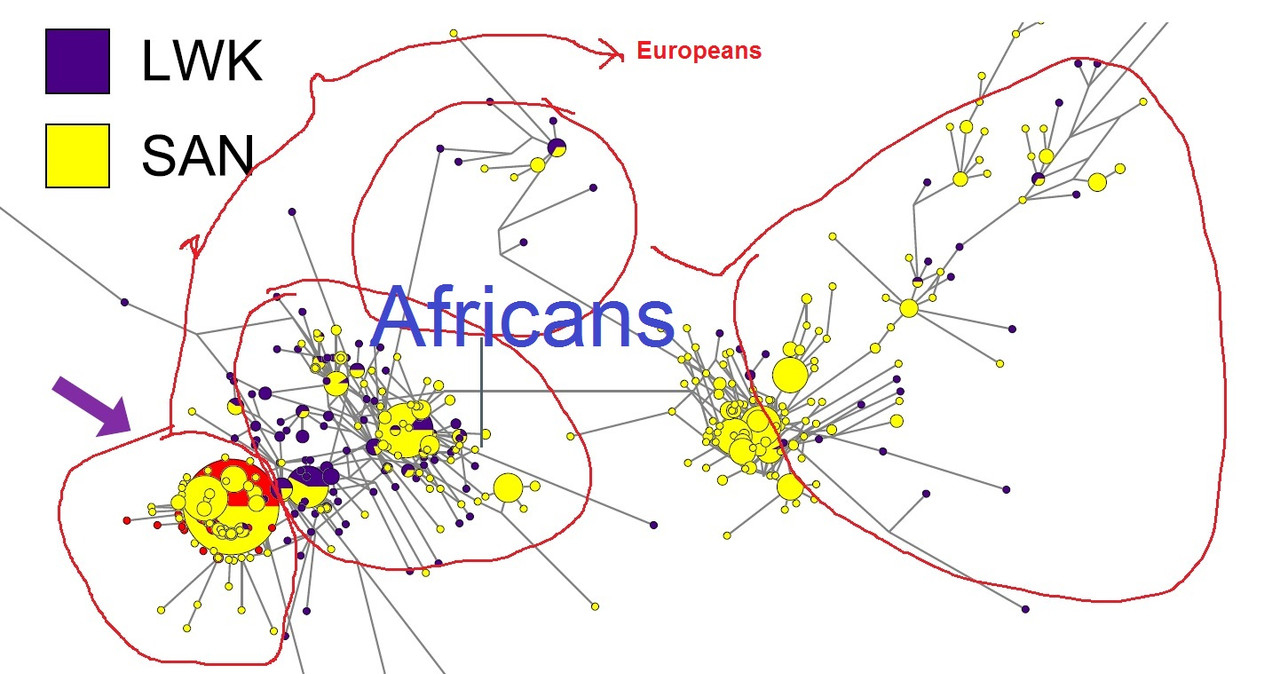
“Haplotype Network of SLC24A5.
We examined the haplotype structure at SLC24A5 to determine whether the derived allele is
identical by descent among populations. While historically isolated, the KhoeSan have received recent gene flow from other
populations, including pastoralists from eastern Africa about 2 kya, Bantu-speaking agropastoralists from eastern Africa
around 500 y ago, and Europeans (including Dutch, French, and German immigrants) about 300 y ago during the colonial era.
The ≠Khomani San and Nama genomes we investigated derive, on average, 12% and 17%, respectively, of their ancestry from Europeans
and 13% and 2%, respectively, from Bantu speakers (the remaining ancestry is primarily KhoeSan; Fig. 2). We obtained **comparative**
SLC24A5 haplotypes from 99 individuals with Northern and Western European ancestry (CEU) and 99 Luhya Bantu speakers from Kenya (LWK) from the 1000 Genomes Project to represent potential source populations for the derived allele. A median-joining
haplotype network based on all individuals exhibits extreme variation across the ancestral haplotypes, which are particularly diverse among
the KhoeSan (Fig. 1B). However, haplotypes with the derived allele are primarily clustered together (SI Appendix, Fig. S1).”
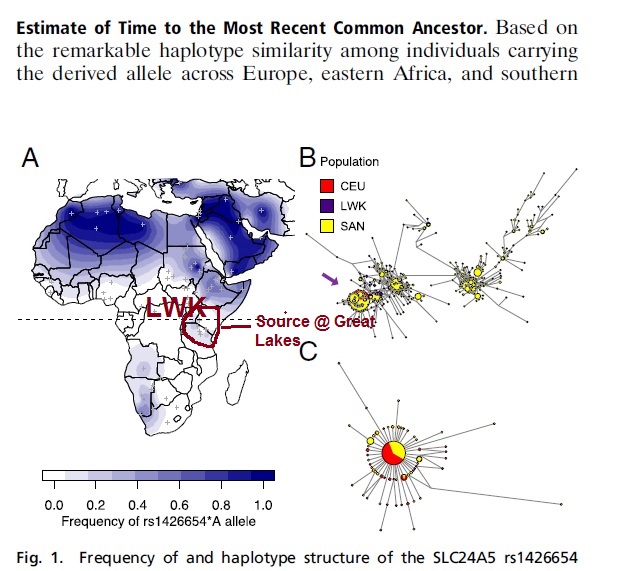
“Estimate of Time to the Most Recent Common Ancestor.
Based on the remarkable haplotype similarity among individuals carrying
the derived allele across Europe, eastern Africa, and southern Africa, we estimated the age of the derived allele within each of these populations. We dated the time to the most recent common
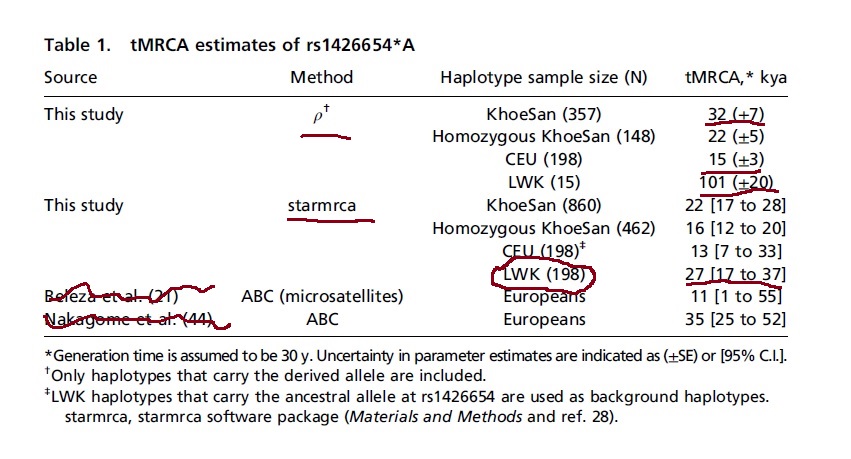
ancestor (tMRCA) of rs1426654*A via **two different methods** (Table 1). First, we calculated the tMRCA from the haplotype
network-based ρ statistic (Materials and Method). We estimated an age of 32 kya (SE is ±7 kya) for all derived haplotypes in KhoeSan.
We then restricted the analysis to haplotypes from homozygous derived individuals to avoid possible phase error, which can artificially
increase allele age estimates by integrating ancestral sequence onto derived haplotypes. This approach resulted in a more recent
tMRCA of 23 kya (±5 kya) for rs1426654*A. European haplotypes have a** tMRCA of 15 kya (±3 kya)**. In contrast, the estimate from
the 15 Luhya derived haplotypes was as old as 101 kya (±20 kya).To explicitly account for recombination between the haplotypes
carrying the ancestral allele at rs1426654 (background haplotypes) and derived haplotypes, we adopted a hidden Markov model that
takes local recombination into account (SI Appendix, Fig. S3) and is valid for selective sweeps. Using both derived and background
haplotypes in KhoeSan, we estimated a tMRCA of 22 kya (95% CI: 17 kya to 28 kya) for all individuals and 16 kya (95% CI: 12 kya to
20 kya) for individuals with the AA homozygote. In comparison, Europeans have a slightly younger tMRCA = 13 kya(using background
haplotypes from LWK), although the CI range largely overlaps with the KhoeSan (95% CI: 7 kya to 33 kya). The Luhya
have the oldest tMRCA = 27 kya (95% CI: 17 kya to 37 kya).”

So both methods show the Kenyans LWK has the light skin genotype that is much older than those in Khoi-San and Europe. WTF. Didn’t I tell you all “dat”. He! He! He!
I should be paid for “tis shyte!!!!”
So if I am reading the data correct. It looks like the pigmentation for light skin always existed in Africa since it was the “ancestral state” for humanoids. It is dated at over 100,000years old in LWK. PRIOR to OOA? Now it looks like black humans first exited to Eurasia. That is why all Europeans were black up to about 5000bc. Not sure about East Asians. Then the EurAfricans rose to dominance in the Great Lakes region and head North, South(Africa) and North East to South Asia.
-----------------
Rapid evolution of a skin-lightening allele in southern African KhoeSan - Meng Lina, Carlos D. Bustamante and Brenna M. Henna
Quote:
“e.g., ∼50% lighter than equatorial Ghanaians. The KhoeSan form the earliest divergence among modern human lineages, branching off from other populations as early as 100 kya to
150 kya. This raises the possibility that the light to intermediate skin of KhoeSan observed today **reflects the ancestral state of the phenotype** and light skin pigmentation in Eurasians
evolved from standing genetic variation within southern Africa .”
“SLC24A5 is among the top signals in positive selection scans of Europeans and Ethiopians, and has long been recognized to play a role in pigmentation pathways.
The nonsynonymous p.Ala111Thr mutation (G → A) at rs1426654 causes a melanin-reduced phenotype in zebrafish. In humans,
the derived allele has swept to fixation or is present at high frequency in many European, Near Eastern, and South Asian populations ; rs1426654 has also been associated with skin pigmentation differences in admixed populations who have recent European ancestors
(i.e., African Americans) and within South Asians. In contrast, the absence of the derived allele in East Asians and
MOST??? sub-Saharan African populations led to the hypothesis that this mutation originated in Europe, or, potentially, the Near East within
the past 10 kya to 35 kya (Table 1).”
“Among the 430 individuals, the derived allele (rs1426654*A) is present at a high frequency of 32.5% in ≠Khomani
And 53.5% in Nama. Queries of rs1426654 in publicly available datasets show that the allele is absent or [/b]at a very low frequency[/b] in
other populations from western, central, and southern Africa (Fig. 1A and SI Appendix, Table S1). Other populations with a moderate to
high frequency of the allele outside Europe have all experienced gene flow from the Near East or Europe during the Holocene, including
those in northern and eastern Africa, and South Asia”
“Haplotype Network of SLC24A5.
We examined the haplotype structure at SLC24A5 to determine whether the derived allele is
identical by descent among populations. While historically isolated, the KhoeSan have received recent gene flow from other
populations, including pastoralists from eastern Africa about 2 kya, Bantu-speaking agropastoralists from eastern Africa
around 500 y ago, and Europeans (including Dutch, French, and German immigrants) about 300 y ago during the colonial era.
The ≠Khomani San and Nama genomes we investigated derive, on average, 12% and 17%, respectively, of their ancestry from Europeans
and 13% and 2%, respectively, from Bantu speakers (the remaining ancestry is primarily KhoeSan; Fig. 2). We obtained **comparative**
SLC24A5 haplotypes from 99 individuals with Northern and Western European ancestry (CEU) and 99 Luhya Bantu speakers from Kenya (LWK) from the 1000 Genomes Project to represent potential source populations for the derived allele. A median-joining
haplotype network based on all individuals exhibits extreme variation across the ancestral haplotypes, which are particularly diverse among
the KhoeSan (Fig. 1B). However, haplotypes with the derived allele are primarily clustered together (SI Appendix, Fig. S1).”
“Estimate of Time to the Most Recent Common Ancestor.
Based on the remarkable haplotype similarity among individuals carrying
the derived allele across Europe, eastern Africa, and southern Africa, we estimated the age of the derived allele within each of these populations. We dated the time to the most recent common
ancestor (tMRCA) of rs1426654*A via **two different methods** (Table 1). First, we calculated the tMRCA from the haplotype
network-based ρ statistic (Materials and Method). We estimated an age of 32 kya (SE is ±7 kya) for all derived haplotypes in KhoeSan.
We then restricted the analysis to haplotypes from homozygous derived individuals to avoid possible phase error, which can artificially
increase allele age estimates by integrating ancestral sequence onto derived haplotypes. This approach resulted in a more recent
tMRCA of 23 kya (±5 kya) for rs1426654*A. European haplotypes have a** tMRCA of 15 kya (±3 kya)**. In contrast, the estimate from
the 15 Luhya derived haplotypes was as old as 101 kya (±20 kya).To explicitly account for recombination between the haplotypes
carrying the ancestral allele at rs1426654 (background haplotypes) and derived haplotypes, we adopted a hidden Markov model that
takes local recombination into account (SI Appendix, Fig. S3) and is valid for selective sweeps. Using both derived and background
haplotypes in KhoeSan, we estimated a tMRCA of 22 kya (95% CI: 17 kya to 28 kya) for all individuals and 16 kya (95% CI: 12 kya to
20 kya) for individuals with the AA homozygote. In comparison, Europeans have a slightly younger tMRCA = 13 kya(using background
haplotypes from LWK), although the CI range largely overlaps with the KhoeSan (95% CI: 7 kya to 33 kya). The Luhya
have the oldest tMRCA = 27 kya (95% CI: 17 kya to 37 kya).”


So both methods show the Kenyans LWK has the light skin genotype that is much older than those in Khoi-San and Europe. WTF. Didn’t I tell you all “dat”. He! He! He!
I should be paid for “tis shyte!!!!”
So if I am reading the data correct. It looks like the pigmentation for light skin always existed in Africa since it was the “ancestral state” for humanoids. It is dated at over 100,000years old in LWK. PRIOR to OOA? Now it looks like black humans first exited to Eurasia. That is why all Europeans were black up to about 5000bc. Not sure about East Asians. Then the EurAfricans rose to dominance in the Great Lakes region and head North, South(Africa) and North East to South Asia.