|
|
Post by djoser-xyyman on Oct 9, 2018 14:31:50 GMT -5
But anyways. I have obtained the VCF files for the Abusirs. I hope to use the VCF files to obtain the CODIS STR via LobSTR. Prior to this I tried Bedtools, BEDOPS and RepeatSeq. All these software seems promising. As I said time to do our own analysis….and publish?. Being banned from ES gave me the time and focus I needed. This is proving to be tough because you need a Linux/Unix background for installing/running these software. These libraries are a pain...getting them right. I am familiar with C++, VBA and BASIC(yes, I am showing my age lol! I am not in my 20’s). Anyways, My programming goes to Windows based- It is called bioinformatics. But it is coming together Use BAM-Analysis kit. There is a trick to running it from Windows. Yes, this is Windows based. Here is what happens. You need to select Chromosomes that has the CODIS STRs which are 2, 3, 4, 5, 6, 7, 8, 11, 12, 13, 15, 17, 18, and 20. The Software will run but at the end all temporary files are deleted/disappears and there is an error message with no data being produced at the end. The trick is to copy these temporary files BEFORE it disappears. These temporary Chromosome related files are VCF, SAM and BAM and BAI etc. They will appear and disappear while the program(BAM-ANALYSIS KIT) is running. Copy them very quickly. The program steps through from Chr 2-20 in about 50min. As I said at the end all meaningful files disappear. For each Abusir it takes about 50minutes to run with about 4-7 minutes per Chr. Some Chr is processed within 3 minutes, so be quick. To do this you need to have the command line window eg CygWin64/Linux window open simultaneously with MS Windows eg Win10. BOTH Windows need to be open as your software runs. Once one Chr is complete that data vanishes. You may end up with 14 VCF files and 14 BAM/BAI/SAM files etc each. These should contain the CODIS STR files …I think. Combine the VCF files(Excel) and run either through BEDtools then LobSTR. I believe RepeatSeq or something similar will also work. Maybe someone who is familiar with Unix/Linux has already have this solved. Point? VCFs files can be obtained from Abusir mummies which we can hopefully use to obtain the CODIS-STRS then plug into spsmart.cesga.es/popstr.php?dataSet=strs_localor PopAffl. |
|
|
|
Post by djoser-xyyman on Oct 10, 2018 6:32:30 GMT -5
After doing a deep dive into bioinformatics I now know most Of these “researchers” that publish these papers are NOT historians, archeologist , Anthropologist etc. They are computer scientist!!!!! They are software engineers, statisticians who are using programs to make incorrect inferences deliberately or through pure ignorance . And why do I say ‘deliberate”. Because their experimental design is really atrocious. Really bad scientific work and inferences. “mistakenly” leaving specific populations out and ignoring the obvious. It is really perplexing that these computer scientist are leading the way on re-writing human history Example – not a historian or archeologist.  |
|
|
|
Post by djoser-xyyman on Oct 10, 2018 6:42:51 GMT -5
|
|
|
|
Post by djoser-xyyman on Oct 10, 2018 6:49:40 GMT -5
deleted
|
|
|
|
Post by djoser-xyyman on Oct 10, 2018 7:04:03 GMT -5
I assumed Paabo was another gay…now I know. SMH. Single mother raised a gay boy. Siiigh. Now He is leading a the charged re-writing human history. Now isn’t that a bitch!!!   |
|
|
|
Post by djoser-xyyman on Oct 10, 2018 7:17:48 GMT -5
So there you have it. Everything goes through these guys(Reich Labs/Max Planck). Can’t believe what I just found out! All these papers/studies on human history has to vetted by these homosexual computer scientists. GTFOH!
Man! This is mind boggling.
Anyways back on topic. I needed a breather
|
|
|
|
Post by djoser-xyyman on Oct 16, 2018 14:17:58 GMT -5
Making progress…Can the CODIS STRs be obtained finally. Anyone can make sense of this JK2911Nuc 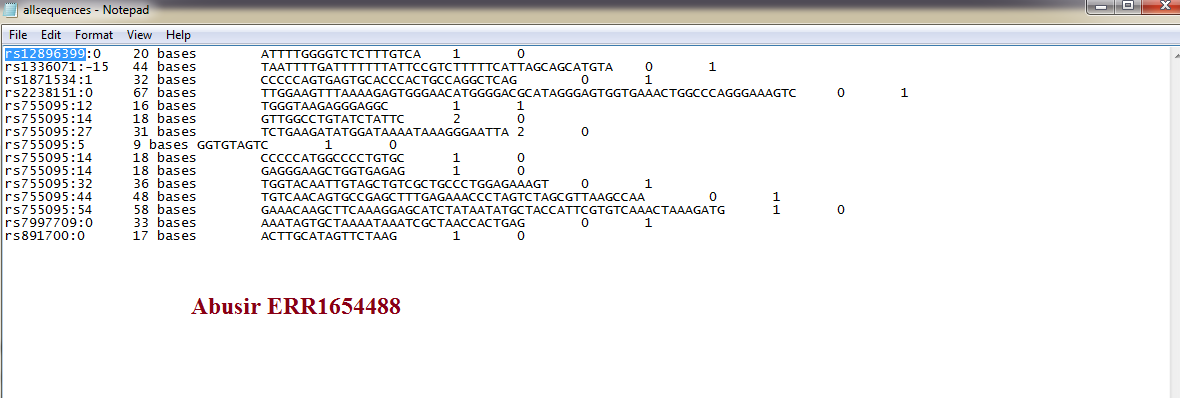 |
|
|
|
Post by kel on Oct 16, 2018 17:40:47 GMT -5
for us non scientists..............please feel free to explain.
|
|
|
|
Post by djoser-xyyman on Oct 17, 2018 10:13:48 GMT -5
This thread is about trying to isolate or pulling CODIS STR from GWA genome wide data(SNPs). Using the Abusir mummies as a test case. Why? Several reasons. 1. The Amarnas (King Tut and his family)were confirmed to closest related to Great Lakes Africans using CODIS STR which was made available by Dr Zink and Hawass through the JAMA Study(2010). We need the same type of data(STR) for the Abusirs. 2. It is impossible for one group(Abusir) of ancient Egyptians who live 50miles away to be European while another group sub-saharan African(Amarnas). IMPOSSIBLE!!!! So Obviously there is data manipulation. 3. Even through their manipulation we have identified the brown component common throughout East Africa(including Khoi-San) also found the ABusirs. Where did this brown component originate? 4. I am using numerous tools(free software) trying to isolate CODIS STRs from the released data files on the Abusir. 5. The above chart is my progress so far. I believe those “rs” is related/located to the CODIS STRs. I need confirmation from the software experts. My programming skills go back to college. I never used it in Industry. But I have done it in college. I am hoping someone reading this can jump in. But I am close. Some software have isolated about 11% SSA ancestry in the Abusir while the released paper said there was NONE. I have some software up and running but there are bugs I am trying to isolate and fix. For starters promising software are HipSTR, Bam Analysis Kit, LobSTR, StRait Razor, BEDtools, glacctools, TreeMix etc 6. An experienced Linux/Unix coder can have this done in an hour!! Not sure why no one is doing it. 7. If you reading my other thread on the gay…Reich, you will understand why this is important. Reich in the interview stated that his models are based upon the assumption that humans remain ISOLATED for 10’s of thousands of years BEFORE admixing. THAT is IMPOSSIBLE. And he knows that. It is deception by Reich and the Max Planck Institute. The genome wide data emerging is consistent with Isolation By Distance and NOT isolation and then re-admixing. Why is that so important……? They are trying to make Africans irrelevant in human and European history. But they know it is a lie and deception. That is why we need to do our own analysis apart from these “barbarians”. That is why he would not perform a DNA test on himself to prove he is a “chosen one”. Afraid to find out the truth and the lies he has been told through his family heritage. 8. The genetic connection between Africa, Southern Europe and even parts of Northern Europe has been establish over 20 years ago. McEvoy, Arnaiz-Villens etc. Reich(Harvard Medical School) and Max Planck is trying to undo that connection established 20 years ago using new approach and models. for us non scientists..............please feel free to explain. |
|
|
|
Post by djoser-xyyman on Oct 17, 2018 10:28:49 GMT -5
I am surprised even Keita did not take that approach. Is he stupid or also on the Euro Team? I am sure he has access to a lot more resources than I. Why would he not employ some newbie research student to do these analysis and publish the results? curoverse.com and other software has shown that Abusir has at least 11% “African” SNPs when the original paper said zero. We in Africana need to start doing our own analysis and posting results. |
|
Deleted
Deleted Member
Posts: 0
|
Post by Deleted on Oct 17, 2018 12:22:44 GMT -5
I am surprised even Keita did not take that approach. Is he stupid or also on the Euro Team? I am sure he has access to a lot more resources than I. Why would he not employ some newbie research student to do these analysis and publish the results? curoverse.com and other software has shown that Abusir has at least 11% “African” SNPs when the original paper said zero. We in Africana need to start doing our own analysis and posting results. Did you try to get in touch with the gentleman with your findings, or are you just a crank spewing venom all around you? |
|
|
|
Post by djoser-xyyman on Oct 17, 2018 13:13:52 GMT -5
BTw – I hope you realize my discourse is not about Atlantis(sorry Kel – just to making a point). But in reality there may have been many huge land masses (Atlantis(s) that is now below the Oceans. Many of the genetic pattern emerging posit that as the only explanation(s)s. May be the Oceanographers got it all wrong. Why? I have many examples. Let us take an easy one….Madagascar. Not only is there a human connection between Africa and Oceanians but there is a flora and fauna connection. So unless the Indonesians WOMEN decided to pack up shop and take the large animals and plants onto boats and decided sail across 4000miles of open Ocean to hook up with black dick....the story is far fetch and too incredulous. It never happened!!!! I came across some old books written by early 19th Century researchers that had doubts. They provided unadultaerated views on what they saw at first hand. There were connecting land masses….Atlantis(s). Same as in West Africa….Cape Verde….Atlantis(s). Here they trying to explain the oriental features of the Malagasy when the Khoi-San up to today carry “oriental” features.   Similar Cape Verde carry R1b-V88 not the Portuguese version of R1b?  |
|
|
|
Post by djoser-xyyman on Oct 17, 2018 13:14:24 GMT -5
An African source of all “Eurasian” populations!!! IBD
Let me explain assuming you don’t understand my point. The Genotype/phenotypic features of Cape Verde and Madagascar can NOT be explained either by Indonesian women sailing to Africa 2000years ago or by Portuguese Sailors co-habiting my West Africans women in the 1600’s. The genetic data do not support these hypotheses.
It is made up BS by Europeans!!!
But I digress. Back on topic
|
|
|
|
Post by djoser-xyyman on Oct 23, 2018 9:17:15 GMT -5
From SAMTools we can pull the STRs since we have the location. Then count the repeats(n). Hopefully they weren't removed prior to being uploaded. 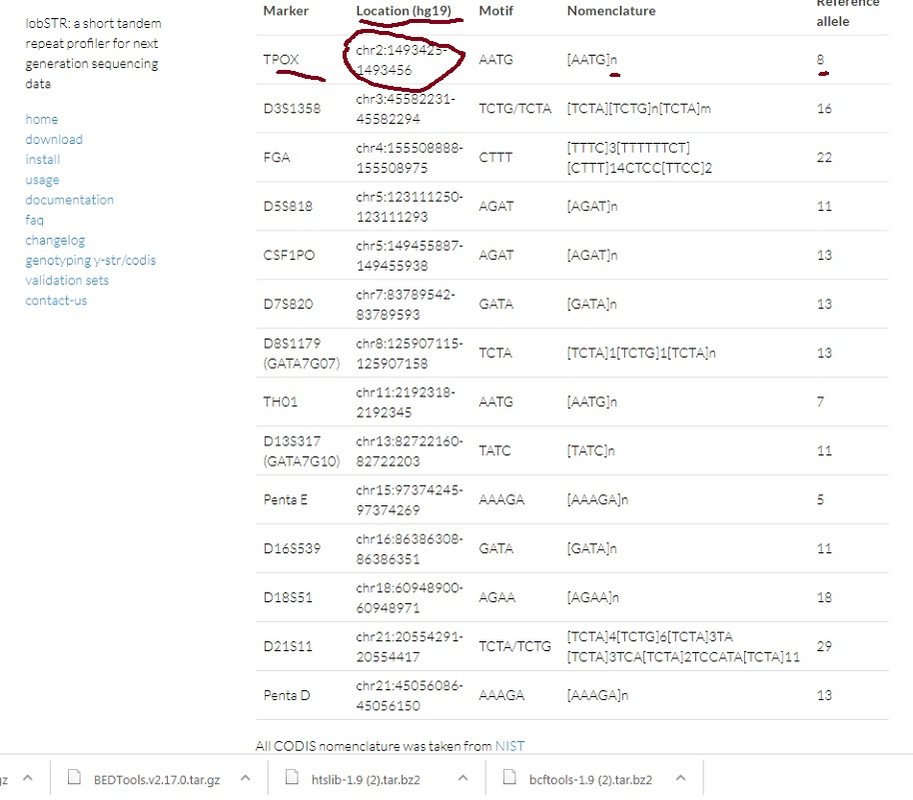 |
|
|
|
Post by truthcentric on Oct 23, 2018 11:56:59 GMT -5
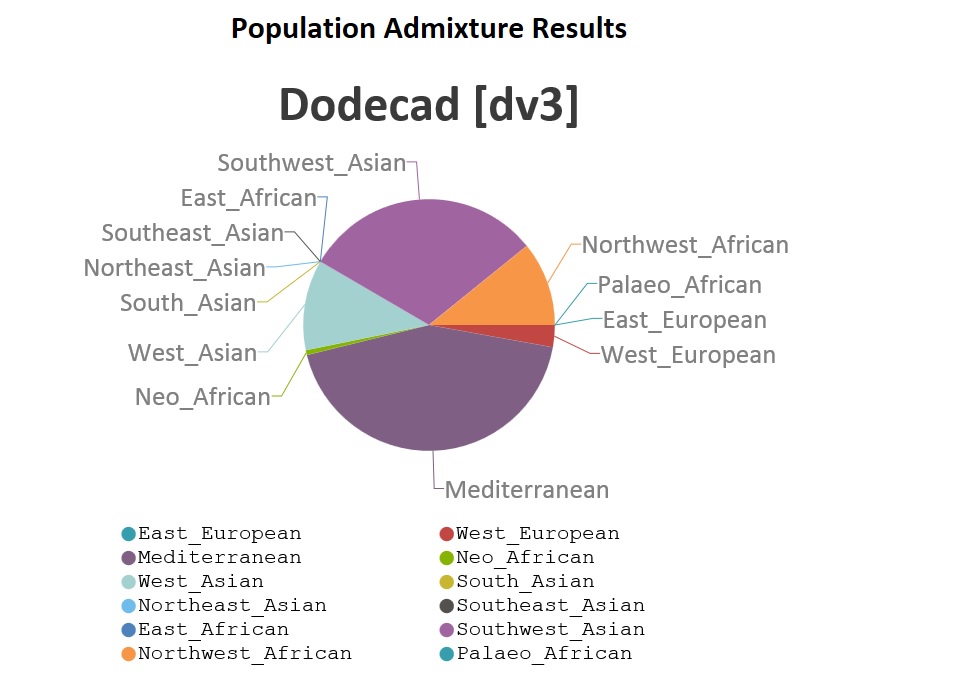 Despite my history of disagreement with the OP author, this is some interesting work done on the Abusir el-Meleq data. I think it would be fascinating to see the same processes (particularly the ones breaking down admixture) applied to the New Kingdom royal mummies. |
|














