|
|
Post by djoser-xyyman on Mar 9, 2019 20:33:02 GMT -5
what is a variant?? The human genome carry about 3 BILLION base pairs. Of those 3 BILLION base pairs about 10 Million are different. ie variants or SNPs. In short of 3,000,000,000 base pairs 3,000,000,000 -10,000,000 =2,990,000,000 bases are the SAME in all humans. We are more similar than different!!!! News flash!! We differ through variants or SNPs. But of the 10,000,000 SNPs only are few are routinely tested in genomic analysis. I chose a larger/different/more random/more transect set of SNPs than Schuenemann. She skewed the results. Plus hse limited SSA to one group of West Africans...the dreaded YRI. Understand how ADMIXTURE works. The intent is to include SNPs that "overlap". So the more populations the more accurate. The more SNPs also the more accurate. She did neither. I think back on ES you said that "back migration" was simply the movement of African variants back into Africa. If so can you expand on this a bit more, such as what sets the border of a genetic variant? |
|
|
|
Post by zarahan on Mar 9, 2019 20:50:28 GMT -5
You asked a good question in a prior post.(oh there was never a movement of humans back into Africa until the Ottoman Turks. There was no movement of variants back into Africa either. That type of gabble-gook comes from ElMeastro) Anyways. To your question. Why is my analysis "better" than Schuenemann? Neither is better. Two different approaches. Schuenemann's is designed for deception. quote from Schuenemann..."The non-UDG and UDG treated libraries were enriched by hybridization to probes targeting approximately 1.24 million genomic SNPs as described previously25. The target SNPs consist of panels 1 and 2 as described in Mathieson et al.41 and Fu et al.26 (see Supplementary Note 2 for details)." What does it mean and what did she do? Schuenemann targeted specific SNPs in the human genome (panels 1 and 2). They were NOT randomly chosen or across the entire human genome. Why? This not rocket science. What I did was took what they gave us(variants/SNP) and compare it with a different global dataset ie population group. I included other populations besides those from HGDP/ I got some from Henn, University of Barcelona and Doron Behar Jews dataset. They showed a totally different outline than Scheuermann's. She stacked the deck to skew the results AWAY from SSA. Abusir are undoubtedly African. ALL OF THEM. Including JK2134. In fact JK2134 carry OLDER VAriants/SNP than the other two that is why it appears so different to the other two. I was surprised to find so much American ancestry in jk2134 but absent in the other two. That pink ancestry is NOT found in Beduoins!!!! Indeed. Stacked decks.. Now where have we seen there before in studies of Africa and the Nile Valley? lol just so the new readers understand the issues at stake.. 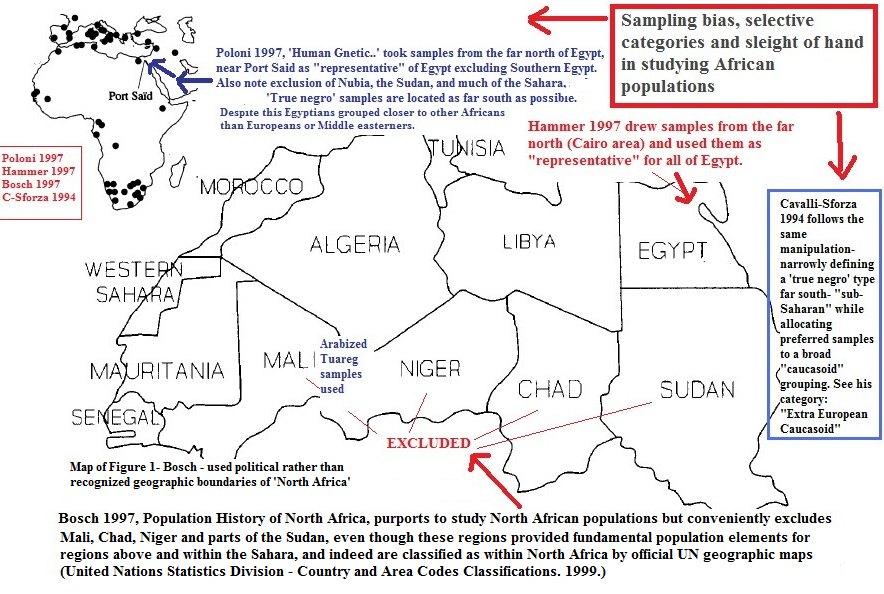  ^^Again, the stacked deck... |
|
|
|
Post by djoser-xyyman on Mar 10, 2019 6:20:54 GMT -5
This file is huge so I needed to cut it down to size. I have to figure out how to upload the entire file. I manually cut this down to size and remove some similar populations. As you can see JK2134 is majority Sub Saharan African. Since it carries a sizeable portion of pink(american) we have to assume it is very old. Keep in mind San and Mbuti carries some "American" ancestry. We see that connection when we look at ADMIXTURE with the Abusir and Skoglund Ancient African. We also see that Bedouins are NOT closely related to Abusir. Yes, one Abusir carry green but the red component negate any European influence or back migration even Levant back-migration. Because this red component is seen on Makrani, and Gond, Paupans and Melanesians....old!!  |
|
|
|
Post by djoser-xyyman on Mar 10, 2019 6:21:55 GMT -5
Including Skoglunds ancient Africans will skew the results further, I will do that next.
|
|
|
|
Post by Tukuler al~Takruri on Sept 18, 2021 12:16:14 GMT -5
|
|
|
|
Post by zarahan on May 22, 2022 21:45:11 GMT -5
Much in this area depends on sampling practices and methods. The high tech CRANID database of skulls for example, draws most of its samples from areas in the further north of Egypt, near the Mediterranean, excluding or downplaying the "darker" historic south, from whence the Dynasties sprung, as some mainstream Egyptologists like Kemp show. The result is a very skewed picture of the population of ancient Egypt. egyptsearchreloaded.proboards.com/post/21734 |
|
|
|
Post by zarahan on May 25, 2022 5:46:47 GMT -5
Then there are skewed DNA games being played- something going on for a long time in the field, since the 1990s or even maybe before.  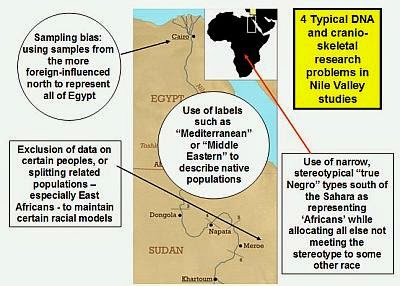 |
|