|
|
Post by djoser-xyyman on Mar 7, 2019 20:44:47 GMT -5
this is a small subset i ran. quick and dirty. 2 Bantu NE, 2 Levant Khazar and 1 Utah White vs 3 Abusir. @ K4. The NE Bantu is present. Doesn't mean much something is up with jk2134 compared to the other 2. Merging with skoglund africans it looks like jk2134 is more older ne african mota like but the other 2 are more southern africans. more to come. Trying to merge this dataset with 16 ancient Africans. getting a parsing error i am trying to work through but yellow is consistent with ibd. with yellow increasing with distance from Africa. interesting to see if skoglund Africans carry these components. I expect they would since jk2911 has all four colors which include the Bantu NE blue and red along with smaller amts of yellow and green. This consistent with yellow and green being African also. I need get more "relatedness" software or back to TreeMix. I ran some TreeMix got results but trying to figure out how to lot and visualize.  |
|
|
|
Post by Tukuler al~Takruri on Mar 7, 2019 21:07:40 GMT -5
Good adjacent continental choices. Utah though mixed is still good for yte Europe. 'Bantu' good enough to rep brwn Africa. Khazar Jew is not a sample. Khazaria was in southeast Europe (Slavo-Turkic) Jews are all over the place. Ditch the politics. Supply the label as provided by the databank it came from please. Which two NE baNtu? (blue n red, important substructure) Which two Jew pops? (all green) =-=-= K=5 is in my previous post. The other 3 K's in the remix    Not sure all the labels are correct. 2 or 3 may be labelling the wrong sample. Please correct where needed Thx |
|
|
|
Post by djoser-xyyman on Mar 7, 2019 21:23:12 GMT -5
Nice work on the Southern vs Northern Africa @k2. I wa strying to have the software recreate the association but I see you did manually.
|
|
|
|
Post by Tukuler al~Takruri on Mar 7, 2019 22:13:52 GMT -5
Hmm. Good. Keep at it.
What do I know.
I thought the pgm's algorithm decided bar graph placement.
But if you can tweak it that'll sidestep ardous supplemental coding.
I look for 100%'s
then majorities
lastly pluralities.
I take geography by
W - E longitude then
S - N latitude
Time depth enters when appropriate.
I think the results allow me to successfully overinterpret ADMIXTURE.
I stare at the color patterns
after aligning them as above so
ancestry mergers and infusions
delineate samples, supersets, and suprasets.
See the African Substructure: Indigenous visavis Migration thread.
Ciao 4 now
|
|
|
|
Post by djoser-xyyman on Mar 7, 2019 23:38:01 GMT -5
yes. Genesis is a "god send"!. Once the phenotype file is created it can be reused. Therefore you can click and drag entire groups using the mouse. no coding necessary. setup the groups in the phenotype file and they can be shifted as a block left or right and even hidden.
|
|
|
|
Post by djoser-xyyman on Mar 9, 2019 8:20:42 GMT -5
quote: " I need get more "relatedness" software or back to TreeMix. I ran some TreeMix got results but trying to figure out how to plot and visualize." got TreeMix up and running with some test files. But I am pilfering some Razib Khan code to save time. He had some errors that I am trying to work through...or reinvent. TreeMix should give relatedness and/or migration edges. quote: "Time depth enters when appropriate."  ?? hmm. Note sure I agree. chronology seems to have a big impact |
|
|
|
Post by Tukuler al~Takruri on Mar 11, 2019 1:15:09 GMT -5
Yes, chronology's importance enters when interpreting first appearance of an ancestry (K color). The exemplar (population with the most of a K color) may not be the originator. Chronology ferrets that out. Chronology can show some K colors in a region arent from the exemplar or the root either. Mt Hora Malawi has K colors associated with Villabruna and Anatolia. Pluralities: Khoe (30%) with the near same amount of San (29%). 'Nuba' * at (20%) is the next substantial part. Atlantic West Africa (8%) and East Africa A (6.5%) are substantial minorities. Even either of the 'Eurasian' contributions (4%; 2.6%) are at like twice that of Mota (1.7%). [*] Nuba (ie, joint NigerCongoKordofanian & NiloSaharan] 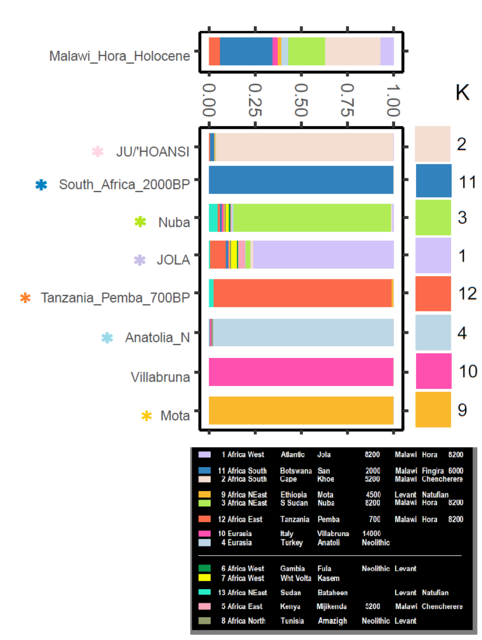 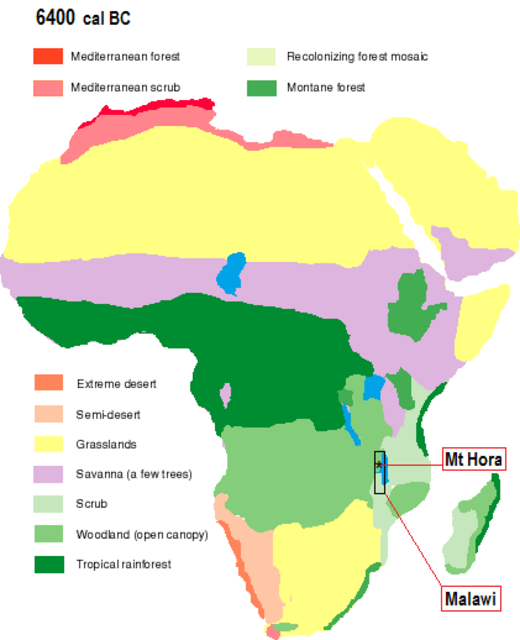 Mt Hora is ~8200 years old and at the West African Monsoon Maximum of the African Humid Period. Southern populations were following fertile savannas and grasslands (their meat & greens markets) north. The 'Sahara' had spouted numerous little lakes, rivers, and marshes from the monsoon rains. Somehow Eurasians (some say their mulattoes and quadroons) supposedly bucked all that northbound traffic. All that fish, game, fruit, and veggies held no appeal. They walked all the 3800 mile way to Malawi (why?) in time enough to be an ancestral component in Mt Hora (infant?) Girl. Amazing. Non-specific West Eurasian is way way down in Malawi just when the Cardium and contemporary cultures were beginning in Europe. |
|
|
|
Post by djoser-xyyman on Mar 11, 2019 12:29:52 GMT -5
for those asking
So what is this telling us?
1. Over 1 million SNPs/Variants were recovered from the ancient Africans
2. Only 135057 SNPs/Variants (over X10) recovered from the Abusir average overlap
3. Since we can only merge variants from “overlapping” datasets that means only 135057 variants will be merged from the Abusir/global/ancient Africans. The Abusir are the rate determining step …ie the bottleneck ie the limitation. The fewest varaiants are from the 3Abusir.
So it seems it is possible for me to merge and perform ADMIXTURE on the 3 Abusir, 16 Ancient Africans and my global dataset of about 2400 modern humans from across the globe
More to come….
--------------------------------------------
plink --bfile skoglund16africans --bmerge abusir-global-fix.bed abusir-global-fix.bim abusir-global-fix.fam --make-bed --out abusir-global-Skog
Warning: Multiple chromosomes seen for variant '.'.
Warning: Multiple chromosomes seen for variant '.'.
Warning: Multiple chromosomes seen for variant '.'.
1233013 markers loaded from skoglund16africans.bim.
135162 markers to be merged from abusir-global-fix.bim.
Of these, 3823 are new, while 131339 are present in the base dataset.
Error: 24407 variants with 3+ alleles present.
-------------------------------------------------
plink --bfile abusir-global-fix --bmerge skoglund16africans.bed skoglund16africans.bim skoglund16africans.fam --make-bed --out abusir-global-Skog2
Working directory: C:\XXXXXX\plink
Start time: Mon Mar 11 13:13:23 2019
Random number seed: 1552324403
8083 MB RAM detected; reserving 4041 MB for main workspace.
2450 people loaded from abusir-global-fix.fam.
16 people to be merged from skoglund16africans.fam.
Of these, 16 are new, while 0 are present in the base dataset.
Warning: Multiple positions seen for variant '.'.
Warning: Multiple positions seen for variant '.'.
Warning: Multiple positions seen for variant '.'.
Warning: Multiple chromosomes seen for variant '.'.
Warning: Multiple chromosomes seen for variant '.'.
Warning: Multiple chromosomes seen for variant '.'.
135057 markers loaded from abusir-global-fix.bim.
1233013 markers to be merged from skoglund16africans.bim.
Of these, 1101779 are new, while 131234 are present in the base dataset.
Error: 24407 variants with 3+ alleles present.
* If you believe this is due to strand inconsistency, try --flip with
abusir-global-Skog2-merge.missnp.
(Warning: if this seems to work, strand errors involving SNPs with A/T or C/G
alleles probably remain in your data. If LD between nearby SNPs is high,
--flip-scan should detect them.)
* If you are dealing with genuine multiallelic variants, we recommend exporting
that subset of the data to VCF (via e.g. '--recode vcf'), merging with
another tool/script, and then importing the result; PLINK is not yet suited
to handling them.
End time: Mon Mar 11 13:13:26 2019
|
|
|
|
Post by djoser-xyyman on Mar 23, 2019 21:23:20 GMT -5
Skoglund 2017 reich.hms.harvard.edu/sites/reich.hms.harvard.edu/files/inline-files/Skoglund_Cell_2017_3.pdfQuote DATA AVAILABILITY Raw sequence data (bam files) from the 15 newly reported ancient individuals is available from the European Nucleotide Archive. The accession number for the sequence data reported in this paper is ENA: PRJEB21878. The newly reported SNP genotyping data is available to researchers who send a signed letter to D.R. containing the following text: ‘‘ (a) I will not distribute the data outside my
collaboration; (b) I will not post the data publicly; (c) I will make no attempt to connect the genetic data to personal identifiers for
the samples; (d) I will use the data only for studies of population history; (e) I will not use the data for any selection studies; (f) I will
not use the data for medical or disease-related analyses; (g) I will not use the data for commercial purposes.’’ reich.hms.harvard.edu/datasets |
|
|
|
Post by djoser-xyyman on Mar 24, 2019 19:09:29 GMT -5
Place holder sourceforge.net/projects/crossmap/files/chain_files/b37tohg19 Skoglund Ancient Africans = GChr37 not hg19(they are very similar) Ancient Iberians = hg19 Abusir = hg19 mydataset - hg19 use LiftOver? quote "Hi, all. I have questions on resource bundles. Are the 'hg19' bundle files just liftover from 'b37' bundles in UCSC-style? If so, why are there some variants in only one version and not the other? For example, the variant 'rs34872315 (on chr1)' is in b37 version of dbsnp137.excluding_sites_after_129.vcf, but not in hg19 version. At first, I thought it's because of the differences in reference genome (vcf files in the bundle are fit for the accompanying reference sequences). But the reference chromosome 1 was the same in both bundles. Can you help me to understand the difference between b37 and hg19 resource bundles?" |
|
|
|
Post by djoser-xyyman on Mar 27, 2019 5:22:02 GMT -5
|
|
|
|
Post by djoser-xyyman on Apr 1, 2019 6:31:05 GMT -5
python3 /home/xxxxx/Crossmap/bin/CrossMap.py
command : python3 /home/xxxxxxx/Crossmap/bin/CrossMap.py vcf GRCh37ToHg19.over.chain.gz I0589.1240k.bam.vcf hg19.fa out-I0589.1240k-bam.vcf
Results
xxxxx@thinkpad:~/genomes/ancientAfricans$ python3 /home/xxxxxxx/Crossmap/bin/CrossMap.py vcf GRCh37ToHg19.over.chain.gz I0589.1240k.bam.vcf hg19.fa out-I0589.1240k-bam.vcf
@ 2019-03-31 18:24:30: Read chain_file: GRCh37ToHg19.over.chain.gz
@ 2019-03-31 18:24:30: Updating contig field ...
@ 2019-03-31 18:24:36: Total entries: 213833
@ 2019-03-31 18:24:36: Failed to map: xxxxxx
xxxxxxxxx@thinkpad:~/genomes/ancientAfricans$
|
|