|
|
Post by clouddesignc7 on Sept 21, 2016 10:51:56 GMT -5
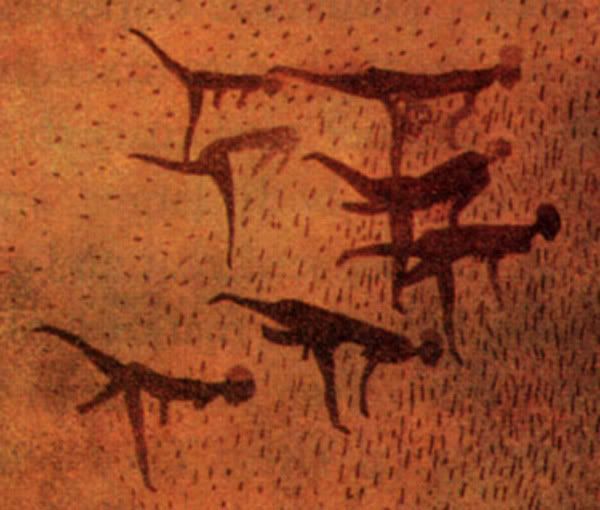
What I quote (also by whatbox) about the Sahara below is based off of the fact that it DID NOT EXIST in pre-pharaohnic times, in the "Mesolithic" times right before Ancient Egypt was started. This is big, I repeat, the Sahara desert as a barrier DID NOT EXIST in those pre-pharaohnic times. (So the Sahara desert as a barrier didn't exist; there were no "sub-Saharan Africans" because sub-Saharan Africans (and animals) (from south of the Sahara) roamed all over Africa. There were just Africans. Notice the giraffe in the Cave painting from Egypt by the way.)
|
|
|
|
Post by clouddesignc7 on Sept 21, 2016 11:09:45 GMT -5
People from far SOUTH roamed far to the NORTH in what are now Arab-Berber areas [/i] [/QB][/QUOTE] [/p][/p][/p][/p][/p] |
|
|
|
Post by clouddesignc7 on Sept 21, 2016 11:24:53 GMT -5
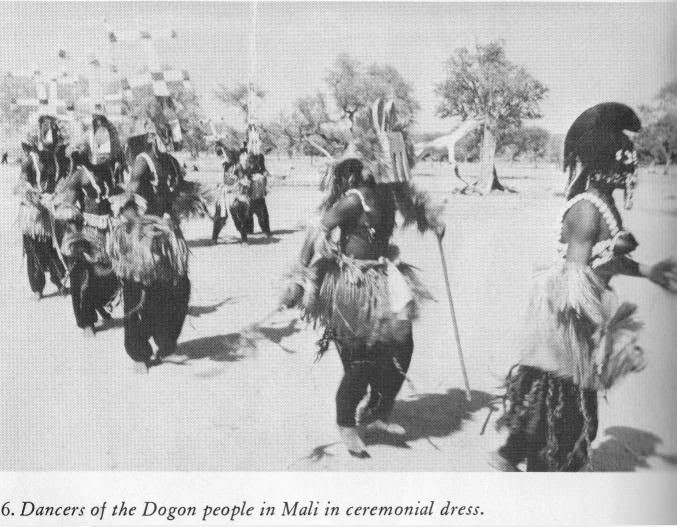
More art from the Sahara
|
|
|
|
Post by clouddesignc7 on Sept 21, 2016 11:34:49 GMT -5
[/list] [/QB][/QUOTE]
|
|
|
|
Post by clouddesignc7 on Sept 21, 2016 12:05:16 GMT -5
E3a / E1b1a / E-M2 found in Egypt!!! and the Middle East (and found to be ancient in Egypt):
This first study is Mitochondrial (that's mtDNA) and doesn't mention M2, it just serves as a prelude to what follows:
Global male inputs from Sub-Saharan Africa and Asia across Iran, not the Levant, into the Arabian Peninsula have been estimated in this study, as 13.4% and 16.6% from both source areas respectively. Recent mtDNA studies on the same Arabian Peninsula countries [7-9,12] have confirmed a notable female-driven sub-Saharan African input with a mean value around 15% for all the Peninsula, although frequencies as high as 60% have been detected in Hadramawt populations of Yemen [9]. Curiously, the Iranian female flow (18%) was also rather similar to that calculated for Africa. Although a slight ratio excess of Sub-Saharan African female versus male gene flow is detected (1.12) we do not found the strong sexual bias proposed by other authors for Arabian populations and attributed to the peculiarities of the recent slave-trade [12,36]. Without dismissing the role mediated by slavery, the geographical distribution of these sub-Saharan African lineages in the Arabian Peninsula seems to indicate a prehistoric entrance of a noticeable portion of these lineages that participated in the building of the primitive Arabian population [8,9]. - Khaled K Abu Amero et. al. 2009, Saudi Arabian Y-Chromosome diversity and its relationship with nearby regions[/color] *** Thoughts about the comedic effect politics may play in all these studies aside: Them saying that a portion of this African gene-flow, even being maternal, is prehistoric and not due to the recent African slave trade is significant, because as interesting as that would be, they're not the only ones to say so. Another study suggests the same, only for paternal lineages: Y-chromosome diversity characterizes the Gulf of Oman
Alicia M Cadenas1, Lev A Zhivotovsky2, Luca L Cavalli-Sforza3, Peter A Underhill3 and Rene J Herrera1
1Department of Biological Sciences, Florida International University, Miami, FL, USA
2N. I. Vavilov Institute of General Genetics, Russian Academy of Sciences, Moscow, Russia
3Department of Genetics, Stanford University, Stanford, CA, USA
Correspondence: Dr RJ Herrera, Department of Biological Sciences, Florida International University, University Park, OE 304, Miami, FL 33199 USA. Tel: +1 305 348 1258; Fax: +1 305 348 1259; E-mail: herrerar@fiu.edu
Received 20 February 2007; Revised 30 August 2007; Accepted 11 September 2007; Published online 10 October 2007.
"Distribution of E3b1-M35 derivatives
The presence of signature sub-Saharan African mtDNA lineages in the south Arabian populations has been attributed to various waves of gene flow to the region, including that associated with the East African slave trade. This is apparent from the exact mtDNA haplotype matches between lineages in Yemen and East Africa, including those associated with the Bantu expansion.20 The presence of the E3a-M2 lineage in Oman (7.4%),4 Yemen (3.2%), UAE (5.5%) and Qatar (2.8%) could lead to the oversimplified conclusion that these chromosomes are also a contribution from the East African slave trade. Mitochondrial DNA analysis of the Yemen Hadramawt indicates recent gene flow (2500 yBP) from Africa to the Arab populations in part through the slave trade, yet an ancient arrival from East Africa is responsible for the Y-chromosome haplotypes." It is also worthy to note that even not all of the recent genetic contribution to the Middle East would have been via slaves. Poor African migrants (and for that matter, any poor people today) may be .. subjugated in the Middle East today (take poor Indian workers in Dubai for instance), but historically this is a major oversimplification. On Egyptian E1b1a (M2) aka Lucotte's "haplogroup 4": Haplotype IV, designating the M2/PN1 subclade, as noted, is found in high frequency in west, central, and sub-equatorial Africa in speakers of Niger-Congo—which may have a special relationship with Nilosaharan—spoken by Nubians; together they might form a superphylum called Kongo-Saharan or Niger-Saharan (see Gregersen 1972, Blench 1995), but this is not fully supported. The spatial distribution of p49a,f TaqI haplotypes in the geographically-widespread speakers of Nilosaharan languages has not been fully characterized, but the notable presence of haplotype IV in Nubians speaking the Eastern Sudanic branch is interesting in that this subgroup is in the Sahelian branch of speakers, whose ancestors may have participated in the domestication of cattle in the eastern Sahara (Ehret 2000, Wendorf and Schild 2001). Sometimes haplotype IV (and the M2 lineage) is seen as being associated with the “Bantu expansion” (2000-3000 bp), but this does not mean that it is not much older, since expansion and origin times cannot be conflated. Haplotype IV has substantial frequencies in upper Egypt and Nubia, greater than VII and VIII, and even V. Bantu languages were never spoken in these regions or Senegal, where M2 is greater than 90 percent in some studies. - Keita, 2005. These genes may not be fully expounded upon or characterized, at least not in any studies I'm aware of, *however*, something else has that has been traced back to West Africa: sickle cell disease. ***
Sickle Cell Anemia in the Middle East and Proximate / Other Regions:The Benin haplotype accounts for HbS associated chromosomes in Sicily,4 Northern Greece,10 Southern Turkey,11 and South West Saudi Arabia,6,7 suggesting that these genes had their origin in West Africa. The Asian haplotype is rarely encountered outside its geographic origin because there have been few large population movements and Indian emigrants have been predominantly from non HbS containing populations. However, it is of interest that the Asian haplotype was first described among descendants of Indian indentured laborers in Jamaica.12
...
From these original foci of the HbS mutation, the gene spread along trading routes to North Africa and the Mediterranean, was transported in large populations to North and South America and the Caribbean during the slave trade, and latterly has spread to Northern Europe by immigration from the Caribbean, directly from Africa to the United Kingdom, France, Belgium, and Holland, and from Turkey to Germany. The relative prevalence of these haplotypes in the Americas reflects the different origins of their African peoples, approximately 70% of HbS associated chromosomes having the Benin haplotype, 10% Senegal and 10% Bantu. Haplotype frequencies in Jamaica are similar to the USA but the Bantu haplotype accounts for the majority of HbS associated chromosomes in Brazil.9
- Graham R. Serjeant, MD, FRCP, MRC Laboratories (Jamaica), University of the West Indies, Kingston.A good map: 
A study on the various types (including Indo-Arabian) in the Middle East (we'll get to how ancient they are after this): Haplotypes of the beta-globin gene as prognostic factors in sickle-cell disease.
el-Hazmi MA, Warsy AS, Bashir N, Beshlawi A, Hussain IR, Temtamy S, Qubaili F.Medical
Biochemistry Department, World Health Organization Collaborating Centre for Haemoglobinopathies, Thalassaemias and Enzymopathies, College of Medicine, King Khalid University Hospital, Riyadh, Saudi Arabia.
The Benin haplotype was found in patients with SEVERE disease, either as homozygous or in combination with another haplotype. The majority of Syrians and Jordanians had the Benin haplotype, and severe disease. However, one in three Syrians and one in five Jordanians had a MILDER disease, and the Saudi-Indian haplotype was identified.
All Saudi patients from south-western and north-western areas, where the disease is generally severe, had the Benin haplotype in the homozygous or heterozygous state. Of the Saudi patients from the eastern area, where a mild form of SCD exists, only 9% had the Benin haplotype. The remainder had the Saudi-Indian haplotype, either in its homozygous or heterozygous state…
Restriction endonuclease restriction sites have provided a useful insight into the normal polymorphic variations in the DNA surrounding various gene loci, where a combination of two or more polymorphic sites has led to the identification of specific haplotype patterns [13,14]. This has been of significance in the study of the regions surrounding the b-globin gene (i.e. the b-globin gene cluster), where several polymorphic sites have been identified, and population differences have been found on analysis of the haplotype pattern [9]. An interesting observation is that the sickle-cell mutation has occurred on chromosomes carrying different polymorphic sites and different b-globin gene haplotypes, and this seems to play a role in the clinical expression of SCD [9].
We compared the haplotype pattern of SCD patients from different Arabic-speaking countries. Benin haplotype was the major haplotype in all countries with a SEVERE presentation of SCD and it was present in both the homozygous and heterozygous state. This was true for those SCD patients from south-western and north-western areas of Saudi Arabia, and for those from Egypt, Jordan and Syrian Arab Republic. On the other hand, patients from the eastern part of Saudi Arabia, who present with a significantly milder clinical picture, carried the Saudi-Indian b-globin gene haplotype either in its homozygous or heterozygous state."
- 1999, Eastern Mediterranean Health JournalModern Middle Easterners would not be the only ones to have severe anemia, PreDynastic Egyptian mummies have it too: Use of the amplification refractory mutation system (ARMS) in the study of HbS in predynastic Egyptian remains.
Dipartimento di Biologia Animale e dell'Uomo, Università degli Studi di Torino.
PMID: 11148985 [PubMed - indexed for MEDLINE]
We conducted a molecular investigation of the presence of sicklemia in six predynastic Egyptian mummies (about 3200 BC) from the Anthropological and Ethnographic Museum of Turin. Previous studies of these remains showed the presence of SEVERE anemia, while histological preparations of mummified tissues revealed hemolytic disorders. DNA was extracted from dental samples with a silica-gel method specific for ancient DNA. A modification of the polymerase chain reaction (PCR), called amplification refractory mutation system (ARMS) was then applied. ARMS is based on specific priming of the PCR and it permits diagnosis of single nucleotide mutations. In this method, amplification can occur only in the presence of the specific mutation being studied. The amplified DNA was analyzed by electrophoresis. In samples of three individuals, there was a band at the level of the HbS mutated fragment, indicating that they were affected by sicklemia. On the basis of our results, we discuss the possible uses of new molecular investigation systems in paleopathological diagnoses of genetic diseases and viral, bacterial and fungal infections.
- Massa et. al., 1999All I have to say to that one is, no wonder! ;D (No wonder about the previous studies attributing M2's presence not to descent / babies of a slave male Middle Eastern Papa, but to ancients) (and fyi male slave input in Mena / Muslim populations seems to have always been negligable if at all |
|
|
|
Post by clouddesignc7 on Sept 21, 2016 12:19:12 GMT -5
A good map:  Notice the Benin sickle cell goes to Egypt and the Middle East. |
|
|
|
Post by clouddesignc7 on Sept 21, 2016 12:32:42 GMT -5
Ramses III


Ramses was found to be E-M2 / E3a / E1b1a (West African)!!
"Genetic kinship analyses revealed identical haplotypes in both mummies (table 1); using the Whit Athey's haplogroup predictor, we determined the Y chromosomal haplogroup E1b1a. The testing of polymorphic autosomal microsatellite loci provided similar results in at least one allele of each marker (table 2)."
- Hawass et al 2012. Revisiing the harem conspiracy and death of Ramesses III. British Medical Journal, BMJ2012;345:e8268




|
|
|
|
Post by anansi on Sept 21, 2016 20:19:10 GMT -5
|
|
|
|
Post by clouddesignc7 on Jan 26, 2017 9:45:03 GMT -5
E-M2 found at 13.9 percent and at 7.7% in Egypt! On Egyptian E1b1a (M2) aka Lucotte's "haplogroup 4": Haplotype IV, designating the M2/PN1 subclade, as noted, is found in high frequency in west, central, and sub-equatorial Africa in speakers of Niger-Congo—which may have a special relationship with Nilosaharan—spoken by Nubians; together they might form a superphylum called Kongo-Saharan or Niger-Saharan (see Gregersen 1972, Blench 1995), but this is not fully supported. The spatial distribution of p49a,f TaqI haplotypes in the geographically-widespread speakers of Nilosaharan languages has not been fully characterized, but the notable presence of haplotype IV in Nubians speaking the Eastern Sudanic branch is interesting in that this subgroup is in the Sahelian branch of speakers, whose ancestors may have participated in the domestication of cattle in the eastern Sahara (Ehret 2000, Wendorf and Schild 2001). Sometimes haplotype IV (and the M2 lineage) is seen as being associated with the “Bantu expansion” (2000-3000 bp), but this does not mean that it is not much older, since expansion and origin times cannot be conflated. Haplotype IV has substantial frequencies in upper Egypt and Nubia, greater than VII and VIII, and even V. Bantu languages were never spoken in these regions or Senegal, where M2 is greater than 90 percent in some studies. - Keita, 2005. E-M2 found at 13.9 percent in a sample size of 274 Egyptians and was found at 7.7 percent in a sample size of 52 Egyptians. So E-M2 was found at 13.9 and 7.7 percent.It was found at 13.9% and 7.7% from that same study: www.academia.edu/10933902/Genetics_Egypt_and_History_Interpreting_Geographical_Patterns_of_Y_Chromosomal_VariationKeita, Genetics, Egypt, and History: Interpreting Geographical Patterns of Y Chromosome variation *** Here's the quote: Haplotype IV, designating the M2/PN1 subclade, as noted, is found in high frequency in west, central, and sub-equatorial Africa in speakers of Niger-Congo—which may have a special relationship with Nilosaharan—spoken by Nubians; together they might form a superphylum called Kongo-Saharan or Niger-Saharan (see Gregersen 1972, Blench 1995), but this is not fully supported. The spatial distribution of p49a,f TaqI haplotypes in the geographically-widespread speakers of Nilosaharan languages has not been fully characterized, but the notable presence of haplotype IV in Nubians speaking the Eastern Sudanic branch is interesting in that this subgroup is in the Sahelian branch of speakers, whose ancestors may have participated in the domestication of cattle in the eastern Sahara (Ehret 2000, Wendorf and Schild 2001). Sometimes haplotype IV (and the M2 lineage) is seen as being associated with the “Bantu expansion” (2000-3000 bp), but this does not mean that it is not much older, since expansion and origin times cannot be conflated. Haplotype IV has substantial frequencies in upper Egypt and Nubia, greater than VII and VIII, and even V. Bantu languages were never spoken in these regions or Senegal, where M2 is greater than 90 percent in some studies.
[...]
Haplotype IV [was found at] 7.7 percent [in] Egypt [...]
Haplotype IV [was found at] 13.9 percent [in] Egypt
|
|
|
|
Post by djoser-xyyman on Jan 26, 2017 10:53:46 GMT -5
Not sure what point you are making by your post?
1. Based Upon RECENT autosomal studies (SNP) modern Egyptians are the MOST admixed with foreign genes than all North Africans. Modern Egyptians are essentially Turks. Sources cited here on ESR. Another main resource have concluded that the invasion of Modern Egypt began about 1300AD(Ottoman invasion). So it was NOT due to invasion from “Arabs” since there is very little difference between indigenous Arabs and some Africans.. Modern Egyptians are heavily admixed with Ottoman Turks. In addition The Amarna’s are essentially sub-saharan Africans based upon autosomal STR. The one male haplogroup that was identified was E-M2 for Rameses III and Man E. Closest genetic match to BOTH Rameses III and Man E …AND the Amarnas were South Africans, Great Lakes Africans and modern West Africans. Modern Levantines and even Magrebians Africans are very distant along with modern Horners.
So I assume the point you are making confirms that yes, Modern Egyptians are essentially foreigners?
Is that the point?
|
|
|
|
Post by asante on Jan 26, 2017 23:35:08 GMT -5
A good map:  Notice the Benin sickle cell goes to Egypt and the Middle East. The issue with that map is that West Africa was not the launching pad of Niger-Congo speakers who characterize the carriers of the adaption. The launching pad was rather the Nile Valley. |
|





 to be given such a title. You have to wonder how ancient Egyptian PHARAOHS got Benin sickle cell -- unless they had some kind now extinct it was the Benin variant, which their spread would explain its spread all over Greece, in Italy, and south-Western Asia in general.
to be given such a title. You have to wonder how ancient Egyptian PHARAOHS got Benin sickle cell -- unless they had some kind now extinct it was the Benin variant, which their spread would explain its spread all over Greece, in Italy, and south-Western Asia in general.













