|
|
Post by djoser-xyyman on May 4, 2018 7:17:32 GMT -5
African geneticist coming into their own? – Botswana Retshabile et al 2018
Whole-Exome Sequencing Reveals Uncaptured Variation and Distinct Ancestry in the Southern African Population of Botswana -
Gaone Retshabile,
collaborative African Genomics Network (CAfGEN) of the H3Africa Consortium
Abstract
Large-scale, population-based genomic studies have provided a context for modern medical genetics. Among such studies, however, African
populations have remained relatively underrepresented. The breadth of genetic diversity across the African continent argues for an
exploration of local genomic context to facilitate burgeoning disease mapping studies in Africa. We sought to characterize genetic variation
and to assess population substructure within a cohort of HIV-positive children from Botswana—a Southern African country that is
regionally underrepresented in genomic databases. Using whole-exome sequencing data from 164 Batswana and comparisons with 150
similarly sequenced HIV-positive Ugandan children, we found that 13%–25% of variation observed among Batswana was not captured
by public databases. Uncaptured variants were significantly enriched (p ¼ 2.2 3 10
16) for coding variants with minor allele frequencies
between 1% and 5% and included predicted-damaging non-synonymous variants. Among variants found in public databases, corresponding
allele frequencies varied widely, with Botswana having significantly higher allele frequencies among rare (<1%) pathogenic
and damaging variants. Batswana clustered with other Southern African populations, but distinctly from 1000 Genomes African populations,
and had limited evidence for admixture with extra-continental ancestries. We also observed a surprising lack of genetic substructure
in Botswana, despite multiple tribal ethnicities and language groups, alongside a higher degree of relatedness than purported
founder populations from the 1000 Genomes project. Our observations reveal a complex, but distinct, ancestral history and genomic
architecture among Batswana and suggest that disease mapping within similar Southern African populations will require a deeper repository
of genetic variation and allelic dependencies than presently exists.
|
|
|
|
Post by djoser-xyyman on May 4, 2018 7:18:13 GMT -5
Anyone? Thoughts?
Quote: “surprising lack of genetic substructure in Botswana, despite multiple tribal ethnicities and language groups, alongside a higher degree of relatedness than purported
founder populations from the 1000 Genomes project. Our observations reveal a complex, but distinct, ancestral history and genomic
architecture among Batswana”
|
|
|
|
Post by djoser-xyyman on May 4, 2018 7:19:10 GMT -5
This I really informative. Notice GWD(Mende?) has a large portion of green component @k4. The green component is lacking in East Africans. Iwo-Eleru?  |
|
|
|
Post by djoser-xyyman on May 4, 2018 7:20:40 GMT -5
Are Africans finally fighting back? Are they going to correct the lies and mis-education?
----
Gaone Retshabile,1 Busisiwe C. Mlotshwa,1 Lesedi Williams,1 Savannah Mwesigwa,2 Gerald Mboowa,2,3
Zhuoyi Huang,4 Navin Rustagi,4 Shanker Swaminathan,5,6 Eric Katagirya,2 Samuel Kyobe,2
Misaki Wayengera,3 Grace P. Kisitu,7 David P. Kateete,2,3 Eddie M. Wampande,2,8 Koketso Maplanka,1
Ishmael Kasvosve,9 Edward D. Pettitt,10 Mogomotsi Matshaba,10,11 Betty Nsangi,7 Marape Marape,10
Masego Tsimako-Johnstone,1 Chester W. Brown,5,12 Fuli Yu,4,5 Adeodata Kekitiinwa,7,11 Moses Joloba,2
Sununguko W. Mpoloka,1 Graeme Mardon,5,13 Gabriel Anabwani,10,11 Neil A. Hanchard,5,6,* and for the
Collaborative African Genomics Network (CAfGEN) of the H3Africa Consortium
|
|
|
|
Post by djoser-xyyman on May 4, 2018 7:21:35 GMT -5
So the Bantu migration never occurred? Didn’t I call it. So they are lying about diseases susceptibility in Africans. Not a surprise.
---
Quotes:
“Given the complex historical ancestry of the Batswana
and the apparent genetic isolation in comparison with
East and West African populations, we then assessed evidence
of structure between our cohort and populations
represented in 1000 Genomes under a maximum likelihood-
based model using ADMIXTURE v.1.3.0 (see Subjects
and Methods). Using 501,963 WES autosomal markers, we
comparedWest African, East African, and Southern African
population groups (Figure 4) from 1000 Genomes and
AGVP. This cross-validation error for the model was minimized
at K ¼ 3 (see Subjects and Methods), with clusters
K2 and K3 separating the populations into West and
East, with Batswana and the other Southern African populations
appearing closer to the East African Luhya (LWK)
population. Cluster K4 distinguished West African populations
and at cluster K5 Batswana and the other Southern
African populations had a component distinct from both
the LWK and the West African populations. Cluster K6
demonstrated the higher level of sub-structure within the
Southern African populations with respect to their West
African and East African counterparts from 1000 Genomes;
this highlights the level of structure that exists between
our populations and the reference populations found
within databases. When we included non-African ancestral
groups (European, South Asian, and East Asian populations)
in the model, we observed minimal??? Eurasian
ancestral components within the Botswana population,
suggesting minimal??? admixture between our cohort and
these population groups (Figure S3).
sharing among Botswana samples alongside our Uganda
cohort and the Finnish population from phase 3 of the
1000 Genomes project (FIN), we found that Botswana had
the longest shared IBD tracks (total and normalized IBD
lengths of 8.84 3 1010 and 1.65 3 106, respectively) of the
three populations, with Uganda also having longer normalized
shared segment lengths (total 3.01 3 1010 and mean
6.74 3 105) than the ****purportedly founder Finnish**** population
of 1000 Genomes (total 8.30 3 109 and mean 4.28 3
105) (Figure 5B). Even after normalizing for sample size
(Subjects and Methods), IBD sharing in Botswana was still
4–5 times higher than that in FIN and was substantially
greater than among the PUR, CLM, FIN, and LWK populations
from 1000 Genomes (Figure 5C), which are known
to have among the smallest effective population sizes
among 1000 Genomes populations1 (see Web Resources
for IBD). IBD sharing in Uganda was closest to the CLM population,
but was still greater than FIN.
identifying
and interpreting such variants in exomes of individuals
with African, and particularly Southern African and
Botswana, ancestry using current iterations of available
public databases will be challenging;1 however, viewed in
the context of our population-level sequence data, many
of the uncaptured variants occurred at frequencies that
would make them less likely to be considered ‘‘damaging’’
or ‘‘deleterious.’’Further, among captured variants, several
uncommon, putatively damaging variants in ClinVar61
and HGMD60 were found to have an appreciably higher
MAF in Botswana, making their pathogenicity more suspect,
at least under a dominant mode of inheritance.
African ancestry has been shown to be positively corre
Thus, at an individual exome
level, current clinical filtering procedures will still result in
multiple candidate variants to be reviewed and validated in
persons of Southern African ancestry. These results bolster
the assertion that the discovery of medically relevant genetic
variants in African populations will likely require
sequenced-based characterization of genetic variation in
the respective, relevant populations”
|
|
|
|
Post by djoser-xyyman on May 4, 2018 7:22:26 GMT -5
Accession Numbers AGVP Datasets: EGAS00001000960/TBA (AGV curated all WGS vcf. files), GAS00001000960/EGAD00001001663 (AGV allele frequencies vcf. files). CAfGEN exome sequence datasets (BAMs and vcfs) are being made publicly available via the European Genome Archive (EGA) in accordance with guidelines agreed upon by the Human Health and Heredity in Africa (H3Africa) Consortium. Supplemental Data Supplemental Data include five figures and five tables and can be found with this article online at doi.org/10.1016/j.ajhg. 2018.03.010. |
|
|
|
Post by djoser-xyyman on May 4, 2018 7:27:00 GMT -5
raw data files, == Web Resources 1000 Genomes, www.internationalgenome.org/ 1000 Genomes IBD Segment Methods, ftp.1000genomes. ebi.ac.uk/vol1/ftp/release/20130502/supporting/ibd_by_pair/ 20150129_IBD_segment_methods.pdf Admixture Manual, www.genetics.ucla.edu/software/ admixture/admixture-manual.pdf APCDR, www.apcdr.org dbNSFP, varianttools.sourceforge.net/Annotation/DbNSFP dbSNP, www.ncbi.nlm.nih.gov/projects/SNP/ Ethnologue, www.ethnologue.com European Genome-phenome Archive (EGA), www.ebi. ac.uk/ega ExAC Browser, exac.broadinstitute.org/ H3Africa, h3africa.org/ |
|
|
|
Post by djoser-xyyman on May 4, 2018 9:06:29 GMT -5
Quote; " We estimated the effective male population size of the African ancestors of Siddis brought to India as ~1,400 individuals" 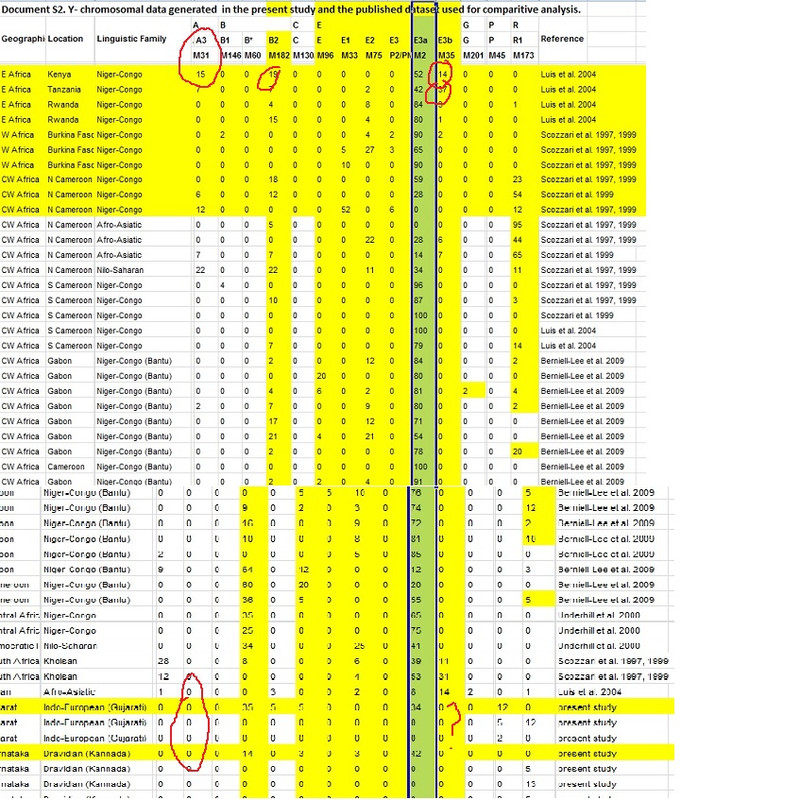 |
|


